Improbable evolution: how life beats the odds

In the 1993 film Jurassic Park, the mathematician character Ian Malcolm, so scene-stealingly played by Jeff Goldblum, rants about science quite a bit, often to poor effect. His references to chaos theory are an utter hash, for example, making it sound indistinguishable from Murphy's Law. And in the speech that summarizes the central conflict of the rampaging dinosaur plot, he insists "life finds a way":
"If there is one thing the history of evolution has taught us, it's that life will not be contained. Life breaks free, expands to new territories, and crashes through barriers, painfully, maybe even dangerously, but, ah, well, there it is."
The florid, unqualified sentiment of Malcolm's quote probably made many biologists in the audience cringe, in keeping with what is described as the character's "deplorable excess of personality." (Again, thank you, Jeff Goldblum!) Malcolm's unwavering fatalism toward biology going rogue may be shared by many opponents of genetic engineering who can never be reassured that their specific fears may be groundless. That's most unfortunate.
Nevertheless, Ian Malcolm wasn't entirely wrong. Time and again, organisms have shown themselves to be adept at evolving around seemingly insurmountable obstacles to their spread and survival. Evolutionary adaptation isn't perfect and inevitable, but it allowed some ancient fish to become four-legged land dwellers, for instance, and some microorganisms to thrive in boiling hot sulfur springs and radioactive pools. Unwavering fatalism about biology going rogue, however, can also breed hysteria over genetic engineering and pandemics.
Biologists have therefore long wanted to understand better the evolutionary mechanisms that enable species to occupy previously forbidden ecological niches -- and the limitations on those mechanisms. Several recent discoveries highlight the importance of that work and provide at least some of the answers.
Fatal flu and viruses that never say die
The most prominent of that work is the much-publicized recent research by independent labs in the Netherlands and Wisconsin on the adaptability of the H5N1 bird flu virus. As I've discussed previously, those scientists discovered that combinations of just five simple mutations in the bird flu's DNA -- which appeared repeatedly during their experiments -- allowed the virus to become a highly infectious, frequently deadly flu in mammals. The work reinforced concerns that the bird flu virus might have the potential to evolve with disturbing ease into the cause of a global pandemic among humans. Indeed, the results were so troubling that flu researchers have elected to impose a temporary moratorium on similar experiments while they consider whether the benefits would compensate for any dangers of the mutant viruses maybe getting loose.
For a pathogen to jump from one host species to another might seem statistically impossible if multiple mutations are needed to effect the transition: if each of those mutations was individually a one-in-a-million shot, the odds of all five occurring spontaneously in one organism would be one in a million trillion trillion (10 to the 30th power). Creationists often make exactly that kind of argument to argue the impossibility of evolution. The flaw in that argument, however, is that it ignores how those mutations can arise as part of a process that makes them collectively more likely.
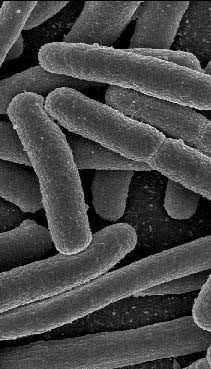
A great insight into exactly that process now comes from a study by Justin R. Meyer and others in the laboratory of Richard E. Lenski of Michigan State University, as published recently in Science. They found a clever way to investigate how organisms can leap past obstacles to their survival by looking at coevolution in the lab between the common bacterium E. coli and the lambda virus that infects it.
Ordinarily, the lambda virus grabs onto a particular protein on E. coli's surface with a matching receptor of its own, then uses that attachment to enter the bacterial cell. By growing E. coli under restrictive conditions, however, Meyer and his colleagues created a strain in which that surface protein was almost completely absent, making infection nearly impossible. One might expect that the lambda virus would rapidly die out in that culture.
Instead, the researchers found that in 24 out of 96 cultures, the viruses evolved an unprecedented ability to infect E. coli via a completely different surface protein -- and in just 15 days. Analysis revealed that they managed this transition with just four small mutations.
Part of what makes this adaptation remarkable, as John N. Thompson of the University of California, Santa Cruz, observed in a commentary accompanying Meyer's paper, is that it defies intuitions about how evolution works:
One of the great metaphors of evolutionary biology ... is that populations evolve toward adaptive peaks separated by adaptive valleys. The peaks are combinations of genes that confer high Darwinian fitness on individuals, whereas the valleys are combinations that confer low fitness. But how can a population move from one peak to another, perhaps higher, peak, across an adaptive valley in which gene combinations are presumed to be maladaptive?
In this case, the answer is in how the bacterium and the virus evolved together.
Mutations in synergy
Lenski's group looked in detail at the specific mutations that enabled the switch to the second protein. They only conferred that ability in combination: individually, they didn't have any affinity for the second protein. Rather, they had affinities for the original surface protein.
That was the key. Although most of the new E. coli strain couldn't make the original protein anymore, a very few had random mutations that restored that ability (it offered no survival advantage). The viral mutations were individually advantageous because on their own, they improved or sustained the virus's ability to infect those few precious, vulnerable cells.
Serendipitously, though, the sets of four mutations in concert allowed a virus to latch onto the second protein, which was an even bigger survival advantage because those viruses could infect all the E. coli in the culture. At every point in the evolutionary process, favored mutations brought at least an incremental extra survival advantage. The mutations were never "aimed" at any fundamental transformation in the viruses' way of life. Yet, together, the four mutations created a narrow bridge across the valley connecting one adaptive peak (affinity for the first protein) to a completely different adaptive peak (affinity for the second protein).
The success of the viruses' evolution did not depend solely on the viruses, however. Meyer and his colleagues discovered that only some of the bacterial cells were good hosts for supporting this evolutionary process: some strains consistently promoted the full transformation while others somehow inhibited it. Specifically, some of the resistant E. coli developed their own mutations that their vulnerability to infection through the second surface protein. As the investigators wrote in their Science paper:
This finding suggests a complex interplay between coevolving phage and bacteria, one that depends on the entire community and its diversity.
The lessons of Meyer's work go beyond viruses and bacteria, however. Applying them to the problem of whether H5N1 is likely to mutate from a bird disease into a human threat, we can surmise that there are no strict guarantees about the outcome -- a fact that even the Dutch and Wisconsin studies corroborate. Ron Fouchier's lab in the Netherlands found that it had created a pathogenic version of the virus that could be transmitted through the air and that killed most of the ferrets it infected. Yoshihiro Kawaoka of the University of Wisconsin-Madison has recently clarified that the altered flu virus his group created was air-transmissible but was not actually deadly to ferrets.
On the other hand, it stands to reason that the more opportunity that pathogens have to associate with new hosts of a different species, the more opportunities they will have to get involved in evolutionary processes that will help them jump to that new species.
Prions outside the brain
All the more worrisome, then, that another study published this past week in Sciencereveals that prions -- the odd infectious proteins responsible for mad cow disease (bovine spongiform encephalopathy, or BSE) in cattle, Creutzfeldt-Jakob disease in humans, and a variety of other neurodegenerative conditions -- can lurk in unexpected places, including species where they don't normally belong.
Prions are abnormally folded versions of a short protein called PrP that have toxic effects on neurons. They are not even so marginally alive as viruses could be said to be, but they nevertheless reproduce by inducing the same misfolding in newly made copies of PrP. Usually, prions are very peculiar to one species: even if BSE prions infect the human nervous system, they don't easily cause misfolding in human PrP, for example.
But now Vincent Béringue and colleagues at the French National Institute for Agronomical Research have shown that prions can also be abundant in the spleen, the appendix, the tonsils, and the lymph nodes. Moreover, those organs seem to be able to harbor prions that would normally only infect other species.
Béringue's group genetically engineered mice that carried copies of human PrP, and then infected them with BSE prions. Only 7 percent of the mice showed signs of the BSE prion in their brain, but 65 percent of the mice had the prion in their spleen. The researchers had similar results when they engineered the mice to carry PrP from sheep and infected them with prions native to elk or hamsters.
The first troubling implication of Béringue's work is that sometimes animals (or people) with no signs of neurological disease could be heavy carriers of prions acquired from other species. In the case of humans, they might unwittingly be able to pass along those infections to others through blood transfusions or other procedures that involved an exchange of lymph or tissues.
But prions can also evolve: as Jiali Li of the Scripps Institute in Florida demonstrated in 2009, variations in the abnormal folding of the PrP can appear over time and variants that reproduce more successfully can proliferate faster in animals. Béringue and his colleagues don't say this, but it seems likely that the spleen and other lymphoid organs could become good places for new species-crossing variants of prions to arise. Whatever the permissive conditions are there that allow the alien prions to survive for extended periods, they might also be conducive to the particles' evolutionary process. It's hard to see how that would be good for us human hosts.
Still, it's important not to assume the worst about these situations, either. Viruses, prions, bacteria, and other pathogens aren't trying to become greater threats to mankind: they aren't trying to be anything. Their evolution is purely a consequence of circumstances that best promote their survival. Life may "find a way," but it isn't particular about what that way is.
•
Related reading on SmartPlanet:
"Study shows how swiftly infectious viruses evolve," by Laura Shin. When a virus can't infect its target cell, how long does it take for it to evolve to successfully invade it again? A new study has a frightening answer.
•
Image: Electron micrograph of an influenza virus. (Credit: Cynthia Goldsmith, CDC/NIH)
This post was originally published on Smartplanet.com
